Institute of Oceanology, Chinese Academy of Sciences
Article Information
- LIU Xudong, SU Chen, FENG Jia, LÜ Junping, LIU Qi, NAN Fangru, LIU Guoxiang, XIE Shulian
- Makinoella parva, a new species of the genus Makinoella (Oocystaceae, Trebouxiophyceae, Chlorophyta)
- Journal of Oceanology and Limnology, 41(6): 2391-2402
- http://dx.doi.org/10.1007/s00343-022-2285-9
Article History
- Received Jul. 23, 2022
- accepted in principle Sep. 13, 2022
- accepted for publication Oct. 28, 2022
2 Key Laboratory of Algal Biology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China
Makinoella Okada is a genus of coccoid green algae which dispose of unique morphology. It was characterized by typical 4–16 celled colonies enclosed in mother cell wall, regularly arranged in a flat plane. Tetrads of the cell were arranged in a quadrilateral shape with subterminal connection (Hegewald et al., 1999). Samples of the genus Makinoella can be easily recognized under microscopic observation due to the large colony size and unique cell arrangement. Recently, only one species, the type species M. tosaensis, was recognized in the genus Makinoella. (Okada, 1949). It was record in four countries up to now (Okada, 1949; Hegewald et al., 1999; Hindák and Hindáková, 2010; Zhang et al., 2016).
Makinoella tosaensis was first discovered in Kochi Prefecture in 1949 (Okada, 1949), and has been reported in several other regions of Japan since then (Fukushima, 1956; Yamagishi and Akiyama, 1984). It was initially classified under the family Oocystaceae (Okada, 1949). The classification was also supported by subsequent studies (Bourrelly, 1966). However, Komárek and Fott (1983) proposed that the alga should be transferred to the family Scenedesmaceae because of the flat 4-celled morphology of the colonies. Due to the similar colony arrangement with Crucigenia Lemmermann, the genus Makinoella was further included in the subfamily Crucigenioideae (Scenedesmaceae) (Komárek and Fott, 1983). Komárek and Fott (1983) suggested a possible classification of the genus Makinoella under class Xanthophyceae due to the presence of numerous chloroplasts and absence of pyrenoids. Between 1996 and 1999, Makinoella was discovered repeatedly at Sookmyung Women's University in Seoul, Republic of Korea, and was recorded by Hegewald et al. in 1999, which was also the first time that the algae were discovered beyond Japan (Hegewald et al., 1999). Hegewald et al. (1999). performed a detailed ultrastructural examination of Makinoella, revealing the perpendicular arrangement of cellulose fibrils adjoining the layer of cell wall. The cell wall ultrastructure unique to the members of family Oocystaceae (Robinson and Preston, 1972; Robinson and White, 1972; Montezinos and Brown, 1978) suggested that the genus Makinoella should be reassigned to family Oocystaceae (Hegewald et al., 1999). In addition, transmission electron microscopy clearly showed that M. tosaensis contains a pyrenoid surrounded by starch sheath (Hegewald et al., 1999; Schnepf and Hegewald, 2000). Subsequently, Hepperle et al. (2000) sequenced the 18S rDNA gene of an algal strain (M. tosaensis SAG28.97) isolated from Korea by Korea by Hegewald et al. (1999) and finally established the taxonomic position of the genus Makinoella via phylogenetic analysis. Accordingly, Makinoella was included in family Oocystaceae (Hepperle et al., 2000) with the closest relationship with Oocystis solitaria. Hindák and Hindáková (2010) collected a strain of Makinoella in the Slovak Republic, which suggested the initial expansion of the genus beyond East Asia. Recently, Zhang et al. (2016) reported two strains from the Wuhan Zoo in Hubei Province and Xishuangbanna tropical botanical garden in Yunnan Province, which was the first record of Makinoella in China.
In this study, we isolated and cultured 17 strains of genus Makinoella from 10 cities of 6 provinces in China. The taxonomic studies of genus Makinoella included morphological observations combined with phylogenetic analysis of multiple molecular markers. The comparison of the ITS-2 secondary structure, together with compensatory base changes (CBCs) and hemi-compensatory base changes (hemi-CBCs), was also used for species separation. The results justified the description of the novel species, M. parva, within the genus.
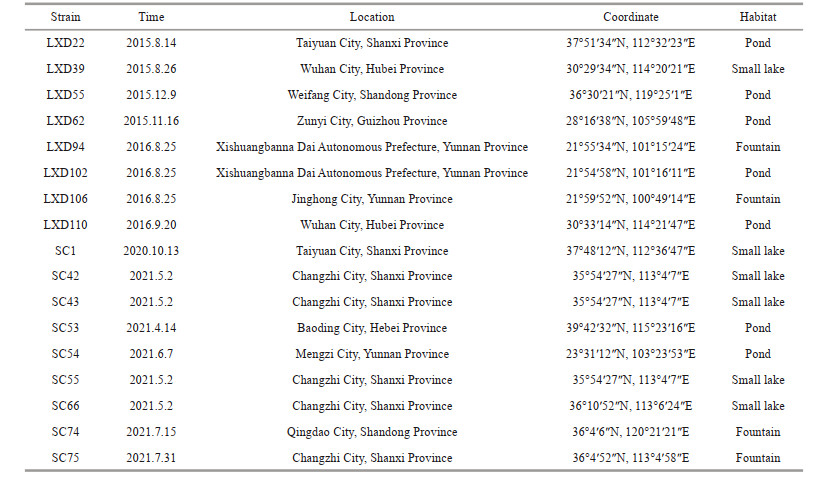
2 MATERIAL AND METHODA total of 17 algal strains were collected in this study (Table 1). Samples were repeatedly retrieved from the water surface with a 10-μm plankton net. The collected samples were partially loaded into 100-mL sampling bottles and transferred to the laboratory. The remaining portions were fixed as a specimen using 10% formalin solution in the field and then preserved in the Algae Research Laboratory of Shanxi University (SXU). A single target cell or colony culture was obtained in the laboratory after multiple isolation and purification steps under the inverted microscope (Olympus CKX41). The pure single-celled algal strains were cultured in liquid BG11 medium under a constant temperature of 25 ℃, a light source of 30–50 μmol photons/(m2·s), and a light-dark cycle of 12 h꞉12 h.
The microscopic morphology of all algal strain samples was observed using an Olympus BX53 light microscope (Olympus Corp., Tokyo, Japan) and an Olympus DP80. The sizes of 50 cells per strain were measured using Olympus cellSens software. The mucilage envelope of the algal strain was shown by negative staining with Indian ink.
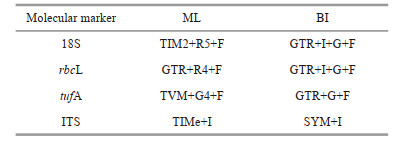
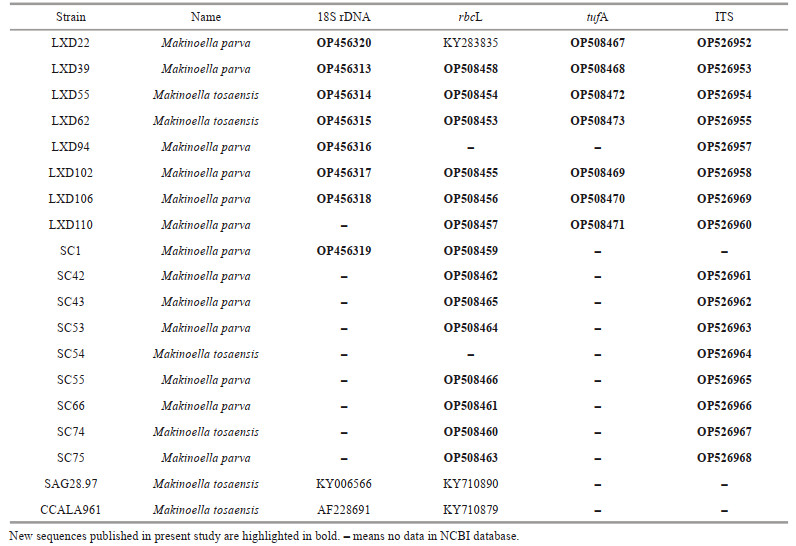
DNA was extracted using a genome extraction kit (Tiangen Biotech Co., China). Four molecular markers, the 18S rDNA, chloroplast genes rbcL and tufA, and ITS region carrying ITS1, 5, 8S rRNA and ITS2, were selected. The primers are shown in Table 2. The amplified product was sent to Beijing Huada Gene Corporation for sequencing. The newly obtained sequences have been submitted to GenBank.
The sequences used for the phylogenetic analysis of 18S rDNA and rbcL were selected based on studies by Štenclová et al. (2017) and Liu et al. (2017, 2018b, 2020). In the absence of tufA and ITS molecular data associated with family Oocystaceae in GenBank, all the sequences used for the phylogenetic analysis based on these two molecular markers were newly obtained in this study. All datasets were aligned using ClustalW (Larkin et al., 2007) and edited manually using Mega 5.2.2 (Tamura et al., 2011). IQ-TREE (Nguyen et al., 2015) and MrBayes 3.2.6 (Ronquist et al., 2012) were used for Maximum Likelihood Estimation (MLE) and Bayesian inference (BI) analysis. The best-fit model was selected by ModelFinder (Kalyaanamoorthy et al., 2017) and listed as Table 3.
The ITS-2 region was detected by ITS2 Database (Schultz et al., 2006). The secondary structure of ITS-2 was constructed based on the RNA structure (Mathews et al., 2004). The hemi-CBCs and CBCs were located by program 4SALE (Seibel et al., 2006, 2008).
3 RESULT 3.1 Morphological observationThe Makinoella parva consisted of a simple colony of 2 to 4 cells, surrounded by the mother cell wall (Fig. 1a & g). Occasionally, solitary cells were observed. The compound colony with multi-generations was seldom formed. The 4-celled colony was composed of two pairs of parallel and symmetrical cells (Fig. 1a & e). The longitudinal axes of the cells were located at an angle (Fig. 1a & i) or even almost perpendicular to each other (Fig. 1c). Occasionally, the four cells were arranged in a square on the same plane (Fig. 1e), and connected via subpolar regions or separated from each other following the expansion of the mother cell wall (Fig. 1a & e). In a 2-celled colony, the cells formed a certain angle or even vertical distribution in different planes (Fig. 1h). The cells were sometimes arranged in the same plane, parallel or non-parallel to each other (Fig. 1d & g). The mother cell wall expanded with wave-like margin (Fig. 1a, c, g, & h) and exhibited obvious bilateral asymmetry in a 2-celled colony (Fig. 1d, g, & h). The surface of mother cell wall showed a fibrillar network structure (Fig. 1d, e, & i) with radially adhering rod-like bacteria (Fig. 1a & c). A thick extracolonial mucilaginous envelope was present (Fig. 1d).

|
| Fig.1 Light microscopy of Makinoella parva and Makinoella tosaensis Microscopic morphology of Makinoella parva in wild habitats (a–h) and under culture conditions (i); a. morphology of typical 4-celled colony; b. morphology of small discoid chloroplasts; c. characteristic morphology of 4 cells arranged vertically in pairs and wavy-like expanded mother cell wall; d. characteristic morphology of the wide colonial mucilaginous envelope; e. characteristic morphology of the broad oval cell in colony; f. characteristic morphology of lamellar chloroplasts; g. morphology of typical 2-celled colony; h. morphology of the 2 cells arranged vertically in asymmetrically expanded mother cell wall; i. characteristic morphology when cultured; microscopic morphology of Makinoella tosaensis in wild habitats (j–k) and under culture conditions (l); j. morphology of typical simple colony; k. morphology of typical compound colony; l. characteristic morphology of the wide colonial mucilaginous envelope. Scale bars: 10 μm. |
Cells were elliptical to broadly elliptical, sometimes slightly asymmetrical (Fig. 1e), and 11.2–21.4 μm long and 7.3–15.2 μm wide. The apical thickening was sometimes visible in older cells. Chloroplasts were discoid or slightly lobated, 4–8 or more, each with a pyrenoid (Fig. 1b, e, & f). The pyrenoid was not always visible (Fig. 1a & d). Numerous metal particles and oil droplets were present in mature cells (Fig. 1c & i). Asexual reproduction involved 2 to 4 autospores. The mother cell wall expanded and then gelatinized or ruptured to release the daughter cells (Fig. 1a).
The Makinoella tosaensis showed similar cell arrangement and colony formation with Makinoella parva. In comparison, the Makinoella tosaensis had more compound colony with 2–3 generations enclosed by old mother cell wall (Fig. 1k). The colony mucilage envelope was thick (Fig. 1l). The cells of Makinoella tosaensis were often broadly elliptical (Fig. 1j), 16–25.6 μm long and 10.6– 19.5 μm wide. When reproductive, the mother cell divided by a transverse wall into 2 autospores (Fig. 1l). The 4 autospores were also common (Fig. 1l). When the daughter cell turned matured, the similar division happened in the mother cell wall. The mother cell wall ruptured to release daughter cells.
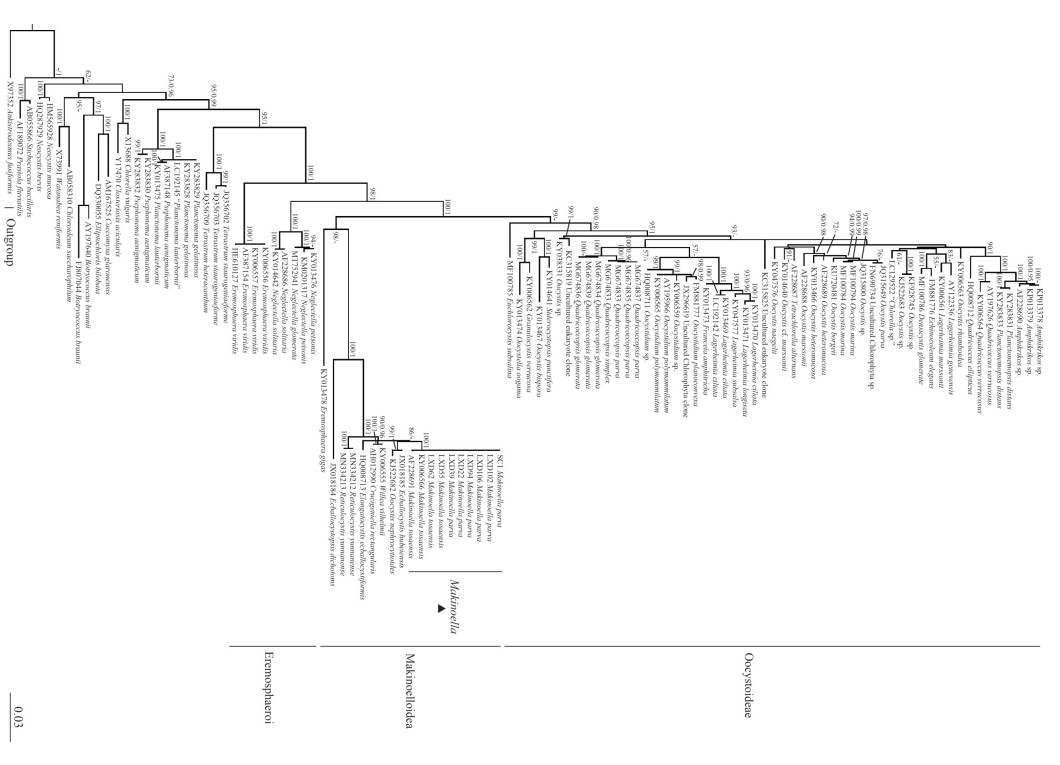
3.2 Phylogenetic analysisIn the present study, 18S rDNA gene was sequenced in eight strains successfully (Table 4), none of which had introns. Phylogenetic analysis was performed in 106 sequences of the representatives of Oocystaceae. Ankistrodesmus fusiformis (Chlorophyta) was selected as the outgroup. Sequence alignment and editing manually yielded a dataset of 1 474 bp. Phylogenetic tree (Fig. 2) of the 18S rDNA showed that the genus Makinoella was included in the subfamily Makinelloideae and was closely related to Ecballocystis hubeiensis and Oocystis nephrocytioides.All the strains of Makinoella clustered on one clade and showed no obviously intra-genus differences.
|
| Fig.2 Phylogenetic tree of Oocystaceae based on 18S rDNA The bootstrap support value of the ML and the posterior probability of the BI are marked at the node. Values above 50 and 0.95 are shown, individually. |
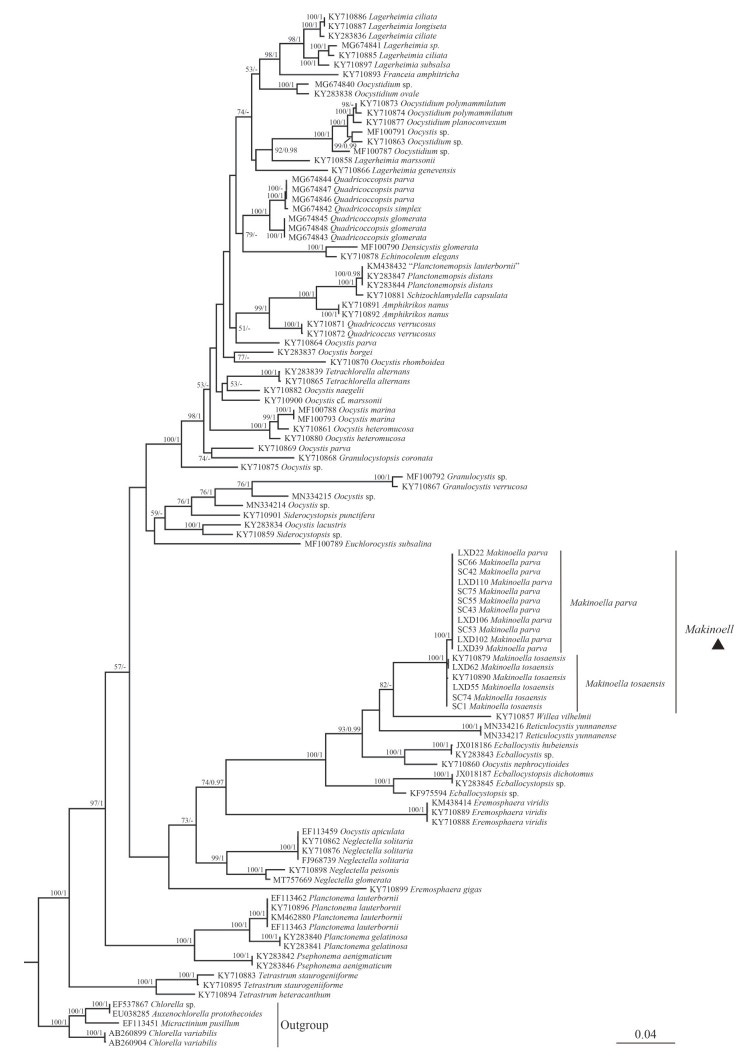
Phylogenetic analyses of the rbcL cpDNA were based on selected 108 representative sequences of the family Oocystaceae including 15 newly obtained sequences of the genus Makinoella in this study. Five strains of Chlorella were selected as the outgroup. The final dataset contained 1 037 bp of rbcL cpDNA. Phylogenetic tree (Fig. 3) showed that the genus Makinoella was clearly divided into two main clades. The basal clade consisted of the Korean strain SAG28.97 (KY710890), the Slovakian strain CCALA961 (KY710879) and our strains LXD55, LXD62, SC1, and SC74. The algal strains LXD22, LXD39, LXD102, LXD106, LXD110, SC42, SC43, SC53, SC55, SC66, and SC75 clustered into the derivative clade with full support.
|
| Fig.3 Phylogenetic tree of Oocystaceae based on rbcL cpDNA The bootstrap support value of the ML and the posterior probability of the BI are marked at the node. Values above 50 and 0.95 are shown, individually. |
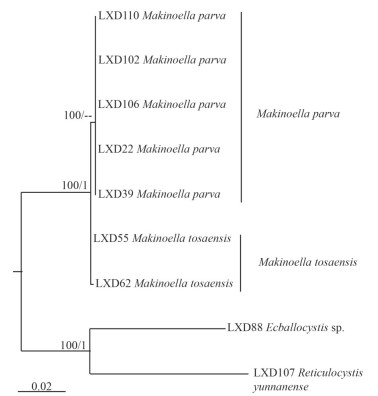
Nine sequences were used for phylogenetic analysis of tufA of the subfamily Makinelloideae, including 7 newly sequenced Makinoella strains and the other two strains, Ecballocystis sp. LXD88 and Reticulocystis yunnanense LXD107, as the outgroup. The sequences were aligned and manually edited in a matrix containing a total of 873 bp of base sites. The phylogenetic tree (Fig. 4) also showed that algal strains LXD22, LXD39, LXD102, LXD106, and LXD110 clustered into one clade, clearly separated from algal strains LXD55 and LXD62 with high support values (100/0.9).
|
| Fig.4 Phylogenetic tree of Makinoella based on tufA cpDNA The bootstrap support value of the ML and the posterior probability of the BI are marked at the node. Values above 50 and 0.95 are shown, individually. |
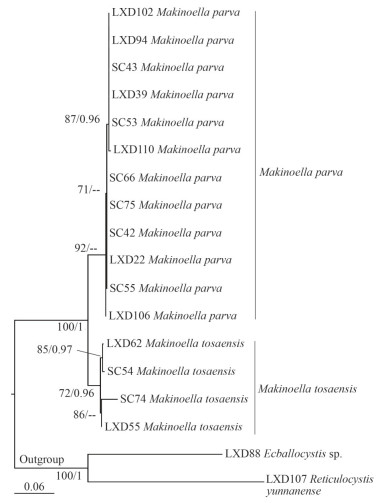
A total of 16 ITS sequences belonging to the genus Makinoella were newly obtained. Sequences of two other representatives of subfamily Makinoelloideae, Ecballocystis sp. LXD88 and Reticulocystis yunnanense LXD107, were used as outgroups. The final matrix contained a total of 620 bp of base sites. The ITS phylogenetic tree (Fig. 5) constructed using MLE and BI analysis showed that the genus Makinoella consisted of two main clades, with strains LXD22, LXD39, LXD94, LXD102, LXD106, LXD110, SC42, SC43, SC53, and SC55 clearly separated from the clade formed by the algal strains LXD55, LXD62, SC54, and SC74, which was also strongly supported statistically (92/0.94).
|
| Fig.5 Phylogenetic tree of Makinoella based on ITS The bootstrap support value of the ML and the posterior probability of the BI are marked at the node. Values above 50 and 0.95 are shown, individually. |
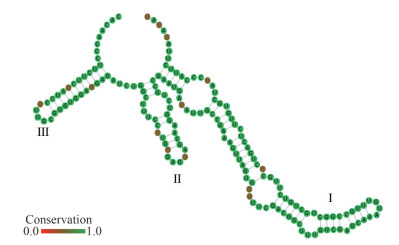
The consensus ITS2 secondary structure model of Makinoella is shown in Fig. 6. Based on the comparison of the ITS2 secondary structure, we found 2 hemi-CBCs in helix Ⅲ and 1 hemi-CBC in helix Ⅱ between the two species.
|
| Fig.6 Consensus ITS-2 secondary structure model of Makinoella Sites with high variability are indicated in red. |
Makinoella is a rare genus of green algae reported only six times worldwide (Okada, 1949; Fukushima, 1956; Yamagishi and Akiyama, 1984; Hegewald et al., 1999; Hindák and Hindáková, 2010; Zhang et al., 2016). In this study, 17 isolated and cultured algal strains (Table 1) suggest that the genus Makinoella has a wide geographical distribution in China. Our findings together with previous studies indicate that the Makinoella species inhabit still waters with small artificial substrates, such as pond, fountains, and small lakes (Okada, 1949; Hegewald et al., 1999; Hindák and Hindáková, 2010; Zhang et al., 2016).
This study entailed a detailed taxonomical investigation of the genus Makinoella, based on morphological observations combined with molecular phylogenetic analysis. The evolutionary relationships within the genus were analyzed using ITS and tufA cpDNA molecular markers for the first time in the family Oocystaceae. Phylogenetic analyses based on ITS, rbcL cpDNA, and tufA cpDNA showed that the genus was clearly divided into two branches with high statistical support, which implied the evolution of at least two distinct taxonomic units in this genus (Figs. 3–5). The CBCs content in the ITS-2 between the branches further supported the separation as two species. Nevertheless, the molecular sequences of the type species have yet to be identified. Therefore, we were unable to classify accurately these two species based on molecular phylogenetic analysis alone.
Morphological comparisons of the two clades of Makinoella strains mentioned above were further conducted. The basal clade consisted of the Korean strain SAG28.97 (Hegewald et al., 1999), the Slovakian strain CCALA961 (Hindák and Hindáková, 2010) and the strains LXD55, LXD62, SC54, and SC74 obtained in this study (Fig. 3) with similar morphology. The morphological characteristics of cells in this clade were similar to the type species, and some of the differences were attributed to not enough detailed observation, which led to wrong original description, and all the strains contained pyrenoids (Okada, 1949). In the family Oocystaceae, the pyrenoid were not always visible, especially when the cells carried a high number of starch grains, oil droplets and other inclusions (Řeháková, 1969), which were typical of M. tosaensis (Hegewald et al., 1999). Therefore, the species of this clade was designated as M. tosaensis, based on the identification of Hegewald et al. (1999) and Hindák and Hindáková (2010).
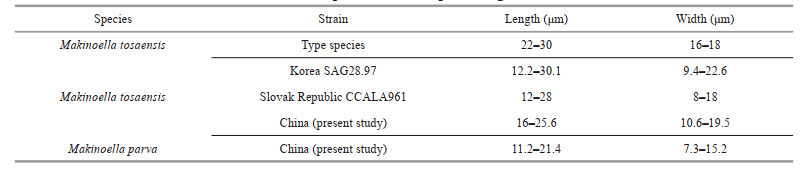
The other clades included newly identified algal strains LXD22, LXD39, LXD94, LXD102, LXD106, LXD110, SC42, SC43, SC53, SC55, SC66, and SC75 (Figs. 3–5), which were significantly smaller in size compared with the type species of Makinoella. The cell size of the type species ((22–30) μm× (16–18) μm) was approximately twice that of the new clades ((11.2–21.4) μm× (7.3–15.2) μm), as shown in Table 5. Cell size was one of the important taxonomic characteristics in the family Oocystaceae (Řeháková, 1969; Komárek and Fott, 1983; Stoyneva et al., 2007), and many species and varieties have been described accordingly, such as Oocystis parva, Quadricoccopsis parva, and Ecballocystis pulvinata var. minor (West and West, 1898; Iyengar, 1932; Liu et al., 2018b). Our results show that the cell size was relatively constant among different strains of the same species (wild or cultured), which was also supported by previous studies of Makinoella reported by Hegewald et al. (1999). Therefore, the cell size was recognized as an effective morphological feature for the classification of the genus Makinoella. Furthermore, the composition of the new clade revealed relatively simpler colonies composed of 2– 4 cells, and rarely included multiple generations. In comparison, the type species consisted of 4–16 cells forming simple or compound colonies. Colony composition was also one of the distinguishing characteristics of the family Oocystaceae on species and generic levels (Komárek and Fott, 1983; Liu et al., 2018b). Based on cell size and colony characteristics, the genus Quadricoccopsis was subdivided into three species: Q. simplex, Q. parva, and Q. glomerata (Liu et al., 2018b). Therefore, based on the smaller cell size, simpler colony composition and independent phylogenetic position, we described the new clade as a new species, M. parva, within the genus Makinoella.
Among the four molecular markers used in this study, 18S rDNA was the most widely used in taxonomical studies of small coccoid green algae. However, the increased data revealed lower variation and phylogenetic signals of 18S rDNA, which interfered with species separation (Bock et al., 2011; Wang et al., 2019). In genus Makinoella, the 18S rDNA also failed to distinguish the different species based on our results. Therefore, distinguishing different genera in Oocystaceae was more appropriate, similar to Scenedesmaceae (Sciuto et al., 2015). The rbcL gene has been associated with Oocystaceae (Liu et al., 2017, 2018a, b, 2020, 2021; Štenclová et al., 2017) and other taxa in Trebouxiophyceae (Song et al., 2015; Sanders et al., 2016). It showed high efficiency for interspecific separation. By contrast, the tufA gene has been recommended for easier amplification and sequencing than the rbcL in the taxonomic study of Chlorophyceae (Vieira et al., 2016), especially in Scenedesmaceae (Sciuto et al., 2015; Wang et al., 2019, 2020). However, in Oocystaceae, the situation was different. Although both are effective for interspecific discrimination of genus Makinoella, the rbcL showed relatively higher amplification efficiency than the tufA. In addition, the rbcL sequences were abundant in the comprehensive analysis by Štenclová et al. (2017), thereby supporting rbcL. In contrast to the two chloroplast genes mentioned above, the ITS molecular markers showed the highest variability. They have been widely used for species separation in Trebouxiophyceae (Krienitz et al., 2004; Bock et al., 2013; Malavasi et al., 2016; Song et al., 2018), especially in Chlorellaceae (Luo et al., 2006, 2010; Krienitz et al., 2010, 2012; Bock et al., 2011). In Oocystaceae, it was also very efficient for the interspecific or intraspecific classification, which was supported by the CBCs and hemi-CBCs results. This study fully supported the application of ITS in the taxonomic study of Oocystaceae. Additional ITS data will be significant for the discovery of species diversity in this family.
Taxonomic assessment
Makinoella parva LIU, SU, LIU et XIE sp. nov. (Fig. 1)
Diagnosis: Colony free-floating, 2–4 cells surrounded by the mother cell wall; cells often not on the same plane or almost perpendicular to each other; mother cell wall wave-like and often with bilateral asymmetry; colonies with thick mucilaginous envelope; cells elliptical to broadly elliptical, 11.2– 21.4 μm long and 7.3–15.2 μm wide; chloroplasts 4–8 or more per cell, discoid or small lamellate, each with a pyrenoid; propagation asexually by 2 or 4 autospores. Species differ from M. tosaensis due to smaller cell size, simpler colony composition and the order of nucleotides in rbcL cpDNA, tufA cpDNA, and ITS.
Type: Baoding City, Hebei Province, China: sample collected in a pond, 39°42′32″N, 115°23′16″E, 14/4/2021, Chen SU (Holotype SXU-SC53). Algae partially illustrated here in light microscopy (Fig. 1).
Reference strain: the living strain was deposited in Freshwater Algae Culture Collection at Institute of Hydrobiology, Wuhan, China as No. FACHB-3564.
Habitat: pond; planktonic; water temperature= 17.6 ℃; pH=8.41.
Etymology: the species is named for the smaller cell size compared with the other species of this genus.
5 CONCLUSIONThe study establishes a new species of the genus Makinoella, M. parva, based on smaller cell volume and simple colony composition and independent phylogenetic position. The rbcL cpDNA and newly used ITS molecular makers showed higher efficiency for interspecific discriminations, and therefore recommended for future taxonomic studies of Oocystaceae.
6 DATA AVAILABILITY STATEMENTThe authors declare that the datasets in this study are available on reasonable request from the corresponding author.
Bock C, Krienitz L, Pröschold T. 2011. Taxonomic reassessment of the genus Chlorella (Trebouxiophyceae) using molecular signatures (barcodes), including description of seven new species. Fottea, 11(2): 293-312.
DOI:10.5507/fot.2011.028 |
Bock C, Luo W, Kusber W H, et al. 2013. Classification of crucigenoid algae: phylogenetic position of the reinstated genus Lemmermannia, Tetrastrum spp. Crucigenia tetrapedia, and C. lauterbornii (Trebouxiophyceae, Chlorophyta). Journal of Phycology, 49(2): 329-339.
DOI:10.1111/jpy.12039 |
Bourrelly P. 1966. Les Algues Deau Douce, Initiation à la Systématique. Tome I: Les Algues Vertes. Ed. N. Boubée, Paris. 512p.
|
Famà P, Wysor B, Kooistra W H C F, et al. 2002. Molecular phylogeny of the genus Caulerpa (Caulerpales, Chlorophyta) inferred from chloroplast tufA gene. Journal of Phycology, 38(5): 1040-1050.
DOI:10.1046/j.1529-8817.2002.t01-1-01237.x |
Fukushima H. 1956. A list of Japanese freshwater algae including the marine species of blue-green algae and fossil diatoms 2. Journal of the Yokohama Municipal University Series C, 46: 1-12.
|
Hegewald E, Schnepf E, Jeon S L. 1999. Report on Makinoella tosaensis Okada (Chlorophyta, Oocystaceae), a species new to Korea. Algae, 14(2): 87-90.
|
Hepperle D, Hegewald E, Krienitz L. 2000. Phylogenetic position of the Oocystaceae (Chlorophyta). Journal of Phycology, 36(3): 590-595.
DOI:10.1046/j.1529-8817.2000.99184.x |
Hindák F, Hindáková A. 2010. First report of Makinoella tosaensis OKADA (Chlorophyta, Chlorococcales, Oocystaceae) outside East Asia. Fottea, 10(1): 141-144.
DOI:10.5507/fot.2010.008 |
Iyengar M O P. 1932. Two little-known genera of green algae (Tetrasporidium and Ecballocystis). Annals of Botany, 46(182): 191-227.
|
Kalyaanamoorthy S, Minh B Q, Wong T K F, et al. 2017. ModelFinder: fast model selection for accurate phylogenetic estimates. Nature Methods, 14(6): 587-589.
DOI:10.1038/nmeth.4285 |
Komárek J, Fott B. 1983. Chlorophyceae (Grünalgen) ordnung: chlorococcales. In: Huber-Pestalozzi G ed. Das Phytoplankton des Süsswassers: Systematik und Biologie. E. Schweizerbart'sche Verlagsbuchhandlung, Stuttgart. p. 1-1044.
|
Krienitz L, Bock C, Kotut K, et al. 2012. Genotypic diversity of Dictyosphaerium-morphospecies (Chlorellaceae, Trebouxiophyceae) in African inland waters, including the description of four new genera. Fottea, 12(2): 231-253.
DOI:10.5507/fot.2012.017 |
Krienitz L, Bock C, Luo W, et al. 2010. Polyphyletic origin of the Dictyosphaerium morphotype within Chlorellaceae (Trebouxiophyceae). Journal of Phycology, 46(3): 559-563.
DOI:10.1111/j.1529-8817.2010.00813.x |
Krienitz L, Hegewald E H, Hepperle D, et al. 2004. Phylogenetic relationship of Chlorella and Parachlorella gen. nov. (Chlorophyta, Trebouxiophyceae). Phycologia, 43(5): 529-542.
DOI:10.2216/i0031-8884-43-5-529.1 |
Larkin M A, Blackshields G, Brown N P, et al. 2007. Clustal W and Clustal X version 2.0. Bioinformatics, 23(21): 2947-2948.
DOI:10.1093/bioinformatics/btm404 |
Liu X D, Wang Q H, Zhu H, et al. 2020. Reticulocystis yunnanense gen. et sp. nov., a new member of freshwater Oocystaceae algae (Trebouxiophyceae, Chlorophyta). European Journal of Phycology, 55(4): 507-516.
DOI:10.1080/09670262.2020.1751303 |
Liu X D, Zhang R, Feng J, et al. 2021. Neglectella glomerata sp. nov., a new species and implications for the systematics of the genus Neglectella (Oocystaceae, Trebouxiophyceae, Chlorophyta). Journal of Oceanology and Limnology, 39(6): 2370-2379.
DOI:10.1007/s00343-020-0356-3 |
Liu X D, Zhu H, Liu B W, et al. 2017. Classification of planctonema-like algae, including a new genus Planctonemopsis gen. nov., a new species Planctonema gelatinosum sp. nov. and a reinstated genus Psephonema (Trebouxiophyceae, Chlorophyta). Journal of Phycology, 53(4): 869-879.
DOI:10.1111/jpy.12551 |
Liu X D, Zhu H, Song H Y, et al. 2018a. Euchlorocystis gen. nov. and Densicystis gen. nov., two new genera of Oocystaceae algae from high-altitude semi-saline habitat (Trebouxiophyceae, Chlorophyta). Journal of Eukaryotic Microbiology, 65(2): 200-210.
DOI:10.1111/jeu.12455 |
Liu X D, Zhu H, Song H Y, et al. 2018b. Quadricoccopsis gen. nov., a new genus of Quadricoccus-like algae in Oocystaceae from China (Trebouxiophyceae, Chlorophyta). Fottea, 18(2): 189-199.
DOI:10.5507/fot.2018.005 |
Luo W, Pflugmacher S, Pröschold T, et al. 2006. Genotype versus phenotype variability in Chlorella and Micractinium (Chlorophyta, Trebouxiophyceae). Protist, 157(3): 315-333.
DOI:10.1016/j.protis.2006.05.006 |
Luo W, Pröschold T, Bock C, et al. 2010. Generic concept in Chlorella-related coccoid green algae (Chlorophyta, Trebouxiophyceae). Plant Biology, 12(3): 545-553.
DOI:10.1111/j.1438-8677.2009.00221.x |
Malavasi V, Škaloud P, Rindi F, et al. 2016. DNA-based taxonomy in ecologically versatile microalgae: a re-evaluation of the species concept within the coccoid green algal genus Coccomyxa (Trebouxiophyceae, Chlorophyta). PLoS One, 11(3): e0151137.
DOI:10.1371/journal.pone.0151137 |
Mathews D H, Disney M D, Childs J L, et al. 2004. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proceedings of the National Academy of Sciences of the United States of America, 101(19): 7287-7292.
DOI:10.1073/pnas.0401799101 |
Montezinos D, Brown R M Jr. 1978. Cell wall biogenesis in Oocystis: experimental alteration of microfibril assembly and orientation. Cytobios, 23(90): 119-139.
|
Nguyen L T, Schmidt H A, Von Haeseler A, et al. 2015. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution, 32(1): 268-274.
DOI:10.1093/molbev/msu300 |
Okada Y. 1949. Makinoella tosaensis, a new genus and species of Oocystaceae. The Journal of Japanese Botany, 24(1-12): 166-168.
|
Řeháková H. 1969. Die Variabilität der Arten der Gattung Oocystis A. Braun. Studies in Phycology. p. 145-196.
|
Robinson D G, Preston R D. 1972. Plasmalemma structure in relation to microfibril biosynthesis in Oocystis. Planta, 104(3): 234-246.
DOI:10.1007/BF00387078 |
Robinson D G, White R K. 1972. The fine structure of Oocystis apiculata W. West with particular reference to the wall. British Phycological Journal, 7(1): 109-118.
DOI:10.1080/00071617200650131 |
Ronquist F, Teslenko M, van der Mark P, et al. 2012. MrBayes 3.2:efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61(3): 539-542.
DOI:10.1093/sysbio/sys029 |
Sanders W B, Pérez-Ortega S, Nelsen M P, et al. 2016. Heveochlorella (Trebouxiophyceae): a little-known genus of unicellular green algae outside the trebouxiales emerges unexpectedly as a major clade of lichen photobionts in foliicolous communities. Journal of Phycology, 52(5): 840-853.
DOI:10.1111/jpy.12446 |
Schnepf E, Hegewald E. 2000. The ultrastructure of Makinoella tosaensis OKADA (Chlorophyta, Oocystaceae). Algological Studies, 97: 79-91.
DOI:10.1127/algol_stud/97/2000/79 |
Schultz J, Müller T, Achtziger M, et al. 2006. The internal transcribed spacer 2 database-a web server for (not only) low level phylogenetic analyses. Nucleic Acids Research, 34(Suppl_2): W704-W707.
DOI:10.1093/nar/gkl129 |
Sciuto K, Lewis L A, Verleyen E, et al. 2015. Chodatodesmus australis sp. nov. (Scenedesmaceae, Chlorophyta) from Antarctica, with the emended description of the genus Chodatodesmus, and circumscription of Flechtneria rotunda gen. et sp. nov.. Journal of Phycology, 51(6): 1172-1188.
DOI:10.1111/jpy.12355 |
Seibel P N, Müller T, Dandekar T, et al. 2006. 4SALE-a tool for synchronous RNA sequence and secondary structure alignment and editing. BMC Bioinformatics, 7: 498.
DOI:10.1186/1471-2105-7-498 |
Seibel P N, Müller T, Dandekar T, et al. 2008. Synchronous visual analysis and editing of RNA sequence and secondary structure alignments using 4SALE. BMC Research Notes, 1: 91.
DOI:10.1186/1756-0500-1-91 |
Song H Y, Liu X D, Hu Y X, et al. 2018. Coronacoccus hengyangensis gen. et sp. nov., a new member of Chlorellaceae (Trebouxiophyceae, Chlorophyta) with radiococcacean morphology. Phycologia, 57(4): 363-373.
DOI:10.2216/17-65.1 |
Song H Y, Zhang Q, Liu G X, et al. 2015. Polulichloris henanensis gen. et sp. nov. (Trebouxiophyceae, Chlorophyta), a novel subaerial coccoid green alga. Phytotaxa, 218(2): 137-146.
DOI:10.11646/phytotaxa.218.2.3 |
Štenclová L, Fučíková K, Kaštovský J, et al. 2017. Molecular and morphological delimitation and generic classification of the family Oocystaceae (Trebouxiophyceae, Chlorophyta). Journal of Phycology, 53(6): 1263-1282.
DOI:10.1111/jpy.12581 |
Stoyneva M P, Cocquyt C, Gärtner G, et al. 2007. Oocystis lacustris CHOD. (Chlorophyta, Trebouxiophyceae) in Lake Tanganyika (Africa). Linzer Biologische Beitrage, 39(1): 571-632.
|
Tamura K, Peterson D, Peterson N, et al. 2011. MEGA5:molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28(10): 2731-2739.
DOI:10.1093/molbev/msr121 |
Vieira H H, Bagatini I L, Guinart C M, et al. 2016. tufA gene as molecular marker for freshwater Chlorophyceae. Algae, 31(2): 155-165.
DOI:10.4490/algae.2016.31.4.14 |
Wang Q H, Liu X D, Li S Y, et al. 2020. Cryptic species inside the genus Hariotina (Scenedesmaceae, Sphaeropleales), with descriptions of four new species in this genus. European Journal of Phycology, 55(4): 373-383.
DOI:10.1080/09670262.2020.1737968 |
Wang Q H, Song H Y, Liu X D, et al. 2019. Morphology and molecular phylogeny of coccoid green algae Coelastrella sensu lato (Scenedesmaceae, Sphaeropeales), including the description of three new species and two new varieties. Journal of Phycology, 55(6): 1290-1305.
DOI:10.1111/jpy.12915 |
West W, West G S. 1898. Notes on freshwater algae. Journal of Botany, British and Foreign, 36: 330-338.
|
Xia S, Zhu H, Cheng Y Y, et al. 2013. Phylogenetic position of Ecballocystis and Ecballocystopsis (Chlorophyta). Fottea, 13(1): 65-75.
DOI:10.5507/fot.2013.006 |
Yamagishi T, Akiyama M. 1984. Photomicrographs of the Freshwater Algae. Uchida Rokakuho Publishing Co., Tokyo. 58p.
|
Zhang Q, Song H Y, Zheng L L, et al. 2016. Makinoella tosaensis Okada, a new recorded genus and species of freshwater Trebouxiophycean algae from China. Journal of Tropical and Subtropical Botany, 24(4): 406-412.
(in Chinese with English abstract) DOI:10.11926/j.issn.1005-3395.2016.04.007 |
 2023, Vol. 41
2023, Vol. 41